For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MAIAPIRIVGDPVLHTPTAPVQVAADGSLPANLNGLISTMYDTMDAAHGVGLAANQIGYGLRVFVYDCAE DCRQTARRRGVVINPILETSEIPETMPDPDTDNEGCLSVPGESFPIGRAQWARVTGLDADGNPVTTEGTG LFARMLQHETGHLDGFLYLDYLIGRHARSAKRAIKSRHWGVPGLSWMPGEVPDPFGP
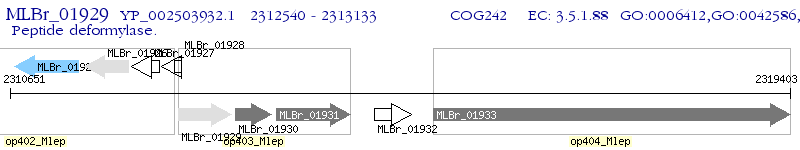
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. leprae Br4923 | MLBr_01929 | def | - | 100% (197) | peptide deformylase |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb0437c | def | 6e-95 | 83.16% (196) | peptide deformylase |
| M. gilvum PYR-GCK | Mflv_0175 | def | 8e-90 | 77.55% (196) | peptide deformylase |
| M. tuberculosis H37Rv | Rv0429c | def | 2e-95 | 83.67% (196) | peptide deformylase |
| M. abscessus ATCC 19977 | MAB_4187 | def | 3e-86 | 77.55% (196) | peptide deformylase |
| M. marinum M | MMAR_0744 | def | 9e-96 | 82.65% (196) | polypeptide deformylase Def |
| M. avium 104 | MAV_4725 | def | 1e-84 | 83.52% (176) | peptide deformylase |
| M. smegmatis MC2 155 | MSMEG_0832 | def | 2e-89 | 77.55% (196) | peptide deformylase |
| M. thermoresistible (build 8) | TH_0951 | def | 4e-91 | 78.57% (196) | PROBABLE POLYPEPTIDE DEFORMYLASE DEF (PDF) (FORMYLMETHIONINE |
| M. ulcerans Agy99 | MUL_1376 | def | 9e-97 | 83.16% (196) | peptide deformylase |
| M. vanbaalenii PYR-1 | Mvan_0732 | def | 2e-92 | 80.61% (196) | peptide deformylase |
CLUSTAL 2.0.9 multiple sequence alignment
MMAR_0744|M.marinum_M MAVVPIRIVGDPVLHTPTSPVPVGADGSLPADLPELIATMYETMDAAHGV
MUL_1376|M.ulcerans_Agy99 MAVVPIRIVGDPVLHTPTSPVPVGADGSLPADLPELIATMYETMDAAHGV
MLBr_01929|M.leprae_Br4923 MAIAPIRIVGDPVLHTPTAPVQVAADGSLPANLNGLISTMYDTMDAAHGV
Mb0437c|M.bovis_AF2122/97 MTVVPIRIVGDPVLHTATTPVTVAADGSLPADLAQLIATMYDTMDAANGV
Rv0429c|M.tuberculosis_H37Rv MAVVPIRIVGDPVLHTATTPVTVAADGSLPADLAQLIATMYDTMDAANGV
MAV_4725|M.avium_104 --------------------MPVGDDGSLPADLVKLIADMYDTMDAAHGV
Mflv_0175|M.gilvum_PYR-GCK MAVRPICIVGDPVLHTATEPIPVGPDGSLPADLADLITDLYDTMDAAHGV
Mvan_0732|M.vanbaalenii_PYR-1 MAVRPIRIVGDPVLHTATEPVPVGADGSLPADLADLITDMYDTMDAAHGV
MSMEG_0832|M.smegmatis_MC2_155 MAVVPIRIVGDPVLHTPTEPVPVGPDGSLPDDLPALIQDMFDTMDAANGV
TH_0951|M.thermoresistible__bu MAVRPIRIVGDPVLHSPTSPVPVGDDGSLPDDLPDLITDMFDTMDAANGV
MAB_4187|M.abscessus_ATCC_1997 MAIVPIRIVGDPVLHTPTQPVPVGPDGSLPDDLPELIANMYETMDAANGV
: *. ***** :* ** :::*****:**
MMAR_0744|M.marinum_M GLAANQIGYGLRLFVYDCADDRGKAAHRRGVVINPVLETSEIPENMPDPD
MUL_1376|M.ulcerans_Agy99 GLAANQIGYGLRLFVYDCADDRRKAAHRRGVVINPVLETSEIPENMPDPD
MLBr_01929|M.leprae_Br4923 GLAANQIGYGLRVFVYDCAEDCRQTARRRGVVINPILETSEIPETMPDPD
Mb0437c|M.bovis_AF2122/97 GLAANQIGCSLRLFVYDCAADRAMTARRRGVVINPVLETSEIPETMPDPD
Rv0429c|M.tuberculosis_H37Rv GLAANQIGCSLRLFVYDCAADRAMTARRRGVVINPVLETSEIPETMPDPD
MAV_4725|M.avium_104 GLAANQIGVGLRVFVYDCADDRGLTERRRGVVVNPVLETSEIPETMPDPD
Mflv_0175|M.gilvum_PYR-GCK GLAANQIGVNKRVFVYDCADARKKTVRRRGVVVNPVLETSEVPETMPDPE
Mvan_0732|M.vanbaalenii_PYR-1 GLAANQIGVSKRVFVYDCADERKKTTRRRGVVINPVLETSEIPETMPDPE
MSMEG_0832|M.smegmatis_MC2_155 GLAANQIGVAKRLFVYDCAPTRGQTTRRRGVVINPVLETSEVPETMPDPD
TH_0951|M.thermoresistible__bu GLAANQIGVSLRVFVYDCAETRGRSTRRRGVVINPVLETSEIPETMPDPD
MAB_4187|M.abscessus_ATCC_1997 GLAANQIGVPLRLFVYDCAETRGGGTRHRGVVINPVLETSEIPETMPDPD
******** *:****** ::****:**:*****:**.****:
MMAR_0744|M.marinum_M NDDEGCLSVPGESFPTGRATWARVTGLDAEGNPVELEGSGLFARMLQHET
MUL_1376|M.ulcerans_Agy99 NDDEGCLSVPGESFPTGRATWARVTGLDAEGNPVELEGSGLFARMLQHET
MLBr_01929|M.leprae_Br4923 TDNEGCLSVPGESFPIGRAQWARVTGLDADGNPVTTEGTGLFARMLQHET
Mb0437c|M.bovis_AF2122/97 TDDEGCLSVPGESFPTGRAKWARVTGLDADGSPVSIEGTGLFARMLQHET
Rv0429c|M.tuberculosis_H37Rv TDDEGCLSVPGESFPTGRAKWARVTGLDADGSPVSIEGTGLFARMLQHET
MAV_4725|M.avium_104 TDDEGCLSVPGESFPTGRASWARVTGLDADGSPVSIEGHGLFARMLQHET
Mflv_0175|M.gilvum_PYR-GCK DDDEGCLSVPGESFPTGRADWARVTGLDADGTPITIEGTDLFARMLQHET
Mvan_0732|M.vanbaalenii_PYR-1 DDDEGCLSVPGESFPTGRADWARVTGLDADGTPITLEGTDLFARMLQHET
MSMEG_0832|M.smegmatis_MC2_155 EDEEGCLSVPGENFPTGRADWARVTGLDADGSPITLEGEDLFARMLQHET
TH_0951|M.thermoresistible__bu TDEEGCLSVPGEQFPTGRADWARVTGLDAEGNPITLEGTDLFARMLQHEV
MAB_4187|M.abscessus_ATCC_1997 DDEEGCLSVPGESFPTGRAGWARVTGLDADGKEVTLEGNDLFARMLQHET
*:*********.** *** *********:*. : ** .*********.
MMAR_0744|M.marinum_M GHLDGYLYLDCLIGRHARSAKRAVKSHGWGVPGLSWLPGEGPDPFGH
MUL_1376|M.ulcerans_Agy99 GHLDGYLYLDCLIGRHARSAKRAVKSHGWGVPGLSWLPGEGPDPFGH
MLBr_01929|M.leprae_Br4923 GHLDGFLYLDYLIGRHARSAKRAIKSRHWGVPGLSWMPGEVPDPFGP
Mb0437c|M.bovis_AF2122/97 GHLDGFLYLDRLIGRYARNAKRAVKSHGWGVPGLSWLPGEDPDPFGH
Rv0429c|M.tuberculosis_H37Rv GHLDGFLYLDRLIGRYARNAKRAVKSHGWGVPGLSWLPGEDPDPFGH
MAV_4725|M.avium_104 GHLDGFLYLDRLIGRYARSAKRAVKSHNWGVPGLSWMPGEGPDPFGH
Mflv_0175|M.gilvum_PYR-GCK GHLDGFLYLDSLIGRNARAAKRAVKSHGWGVPGLTWMPGEDPDPFGH
Mvan_0732|M.vanbaalenii_PYR-1 GHLDGFLYLDRLIGRNARSAKKTVKSHGWGVPGLSWMPGEDPDPFGH
MSMEG_0832|M.smegmatis_MC2_155 GHLDGFLYLDRLVGRYARAAKKAVKRNGWGVPGLSWMPGEVPDPFGH
TH_0951|M.thermoresistible__bu GHLDGYLYLDRLVGRYARAAKRMVKARGWGVPGLSWMPGEVPDPFGH
MAB_4187|M.abscessus_ATCC_1997 GHLDGFLYIDKLIGRNARAAKRAVKSNGWGVPGLSWTPGQVADPFGH
*****:**:* *:** ** **: :* . ******:* **: .****