For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MQRIIGTEVEYGISSPTDPAANPILTSTQAVLAYAAATGIQRAKRTRWDYEVESPLRDARGFDLGRTTGP APIVDADEVGAANMILTNGARLYVDHAHPEYSAPECTNPLDAVIWDKAGERVMEAAARHVASVPGAARLQ LYKNNIDGKGASYGTHENYLMDRQTPFSAIIAGLTPFFVSRQVVTGSGRVGIGPSGDDAGFQLSQRADYI EVEVGLETTLKRGIINTRDEPHADADKYRRLHVIIGDANLAETSTFLKVGTTSLVLDLIESGVDLSDLAL ARPVHAVHVISRDPSLRATVALADGRELTGLALQRIYLDRVAELVSSRDPDPQADQVLQVWAKVLDLLER DPMECADLLDWPAKLRLLEGFRQRENLSWSAPRLHLVDLQYSDVRLDKGLYNRLVARGSMQRMVTEQQVL DAVTRPPSDTRAYFRGECLRKFGADIAAASWDSVIFDLGGDSLVRIPTLEPLRGSKSHVGALLDSVDSAA ELVDQLTT
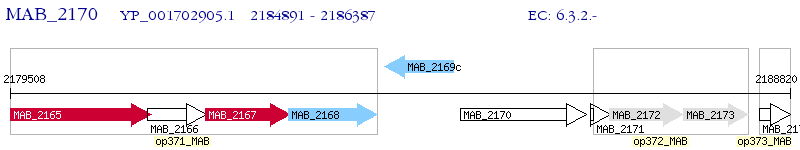
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. abscessus ATCC 19977 | MAB_2170 | - | - | 100% (498) | hypothetical protein MAB_2170 |
| M. abscessus ATCC 19977 | MAB_2183 | - | 3e-80 | 40.24% (415) | hypothetical protein MAB_2183 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb2136c | - | 0.0 | 89.02% (501) | hypothetical protein Mb2136c |
| M. gilvum PYR-GCK | Mflv_3075 | - | 0.0 | 89.42% (501) | hypothetical protein Mflv_3075 |
| M. tuberculosis H37Rv | Rv2112c | - | 0.0 | 89.02% (501) | hypothetical protein Rv2112c |
| M. leprae Br4923 | MLBr_01320 | - | 0.0 | 87.65% (502) | hypothetical protein MLBr_01320 |
| M. marinum M | MMAR_3087 | - | 0.0 | 87.65% (502) | hypothetical protein MMAR_3087 |
| M. avium 104 | MAV_2406 | - | 5e-80 | 40.29% (412) | proteasome component |
| M. smegmatis MC2 155 | MSMEG_3897 | - | 0.0 | 91.35% (497) | proteasome component |
| M. thermoresistible (build 8) | TH_0506 | - | 0.0 | 87.15% (498) | CONSERVED HYPOTHETICAL PROTEIN |
| M. ulcerans Agy99 | MUL_2333 | - | 0.0 | 87.45% (502) | hypothetical protein MUL_2333 |
| M. vanbaalenii PYR-1 | Mvan_3451 | - | 0.0 | 90.64% (502) | hypothetical protein Mvan_3451 |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_3075|M.gilvum_PYR-GCK ------------------------------MSPVGLVTSPSGQTSTLARM
Mvan_3451|M.vanbaalenii_PYR-1 -------------------------------------------------M
MSMEG_3897|M.smegmatis_MC2_155 -------------------------------------------------M
TH_0506|M.thermoresistible__bu --------------------------------------------------
Mb2136c|M.bovis_AF2122/97 MFWVG-------------------GRCLMPASSAARCAARIVGGPRLYGM
Rv2112c|M.tuberculosis_H37Rv MFWVGGPCLMPASSAARCAARIVGGRCLMPASSAARCAARIVGGPRLYGM
MMAR_3087|M.marinum_M -------------------------------------------------M
MUL_2333|M.ulcerans_Agy99 -------------------------------------------------M
MLBr_01320|M.leprae_Br4923 -------------------------------------------------M
MAB_2170|M.abscessus_ATCC_1997 -------------------------------------------------M
MAV_2406|M.avium_104 -------------------------------------------------M
Mflv_3075|M.gilvum_PYR-GCK QR-IIGTEVEYGISSPTDPTANPILTSTQAVLAYAAAAGIQRAKRTRWDY
Mvan_3451|M.vanbaalenii_PYR-1 QR-IIGTEVEYGISSPSDPTANPILTSTQAVLAYAAAAGIQRAKRTRWDY
MSMEG_3897|M.smegmatis_MC2_155 QR-IIGTEVEYGISSPSDPTANPILTSTQAVLAYAAAAGIQRAKRTRWDY
TH_0506|M.thermoresistible__bu ----MGTEVEYGISSPSDPTANPILTSTQAVLAYAAAAGLQRGKRTRWDY
Mb2136c|M.bovis_AF2122/97 QR-IIGTEVEYGISSPSDPTANPILTSTQAVLAYAAAAGIQRAKRTRWDY
Rv2112c|M.tuberculosis_H37Rv QR-IIGTEVEYGISSPSDPTANPILTSTQAVLAYAAAAGIQRAKRTRWDY
MMAR_3087|M.marinum_M QR-IIGTEVEYGISSPSDPTANPILTSTQAVLAYAAAAGIQRAKRTRWDY
MUL_2333|M.ulcerans_Agy99 QR-IIGTEVEYGISSPSDPTANPILTSTQAVLAYAAAAGIQRAKRTRWDY
MLBr_01320|M.leprae_Br4923 QR-IIGTEVEYGISSPSDPTANPILTSTQAVLAYAAAAGIQRVKRTRWDY
MAB_2170|M.abscessus_ATCC_1997 QR-IIGTEVEYGISSPTDPAANPILTSTQAVLAYAAATGIQRAKRTRWDY
MAV_2406|M.avium_104 QRRIMGIETEFGVTCT-----------------------FHGHRRLSPDE
:* *.*:*::.. :: :* *
Mflv_3075|M.gilvum_PYR-GCK EVESPLRDARGFDLSRSSGPPPVVDADEVGAANMILTNGARLYVDH-AHP
Mvan_3451|M.vanbaalenii_PYR-1 EVESPLRDARGFDLSRSSGPPPVVDADEVGAANMILTNGARLYVDH-AHP
MSMEG_3897|M.smegmatis_MC2_155 EVESPLRDARGFDLSRSSGPPPIVDADEVGAANMILTNGARLYVDH-AHP
TH_0506|M.thermoresistible__bu EVESPLRDARGFDLSRSAGPPPIVDADEVGAANMILTNGARLYVDH-AHP
Mb2136c|M.bovis_AF2122/97 EVESPLRDARGFDLSRSAGPPPVVDADEVGAANMILTNGARLYVDH-AHP
Rv2112c|M.tuberculosis_H37Rv EVESPLRDARGFDLSRSAGPPPVVDADEVGAANMILTNGARLYVDH-AHP
MMAR_3087|M.marinum_M EVESPLRDARGFDLSRSAGPPPVVDADEVGAANMILTNGARLYVDH-AHP
MUL_2333|M.ulcerans_Agy99 EVESPLRDARGFDLSRSAGPPPVVDADEVGAANMILTNGARLYVDH-AHP
MLBr_01320|M.leprae_Br4923 EVESPLRDARGFDLSRSAGPPPVVDADEVGAANMILTNGARLYVDH-AHP
MAB_2170|M.abscessus_ATCC_1997 EVESPLRDARGFDLGRTTGPAPIVDADEVGAANMILTNGARLYVDH-AHP
MAV_2406|M.avium_104 VARYLFR--RVVSWGRSS--------------NVFLRNGARLYLDVGSHP
.. :* * .. .*:: *::* ******:* :**
Mflv_3075|M.gilvum_PYR-GCK EYSAPEVTDPMDAVIWDKAGERVME----AAARHVASVPGAAKLQLYKNN
Mvan_3451|M.vanbaalenii_PYR-1 EYSAPECTDPMDAVIWDKAGERVME----AAARHVASVPGAAKLQLYKNN
MSMEG_3897|M.smegmatis_MC2_155 EYSAPECTDPMDAVIWDKAGERVME----AAARHVASVPGAAKLQLYKNN
TH_0506|M.thermoresistible__bu EYAAPETTDPLDAVIWDKAGERVME----AAARHVASVPGAVKLQLYKNN
Mb2136c|M.bovis_AF2122/97 EYSAPECTDPLDAVIWDKAGERVME----AAARHVASVPGAAKLQLYKNN
Rv2112c|M.tuberculosis_H37Rv EYSAPECTDPLDAVIWDKAGERVME----AAARHVASVPGAAKLQLYKNN
MMAR_3087|M.marinum_M EYSAPECTDPLDAVIWDKAGERVME----AAARHVASVPGAAKLQLYKNN
MUL_2333|M.ulcerans_Agy99 EYSAPECTDPLDAVIWDKAGERVME----AAARHVASVPGAAKLQLYKNN
MLBr_01320|M.leprae_Br4923 EYSAPECTDPLDAVIWDKAGERVME----AAARHVASVPGAAKLQLYKNN
MAB_2170|M.abscessus_ATCC_1997 EYSAPECTNPLDAVIWDKAGERVME----AAARHVASVPGAARLQLYKNN
MAV_2406|M.avium_104 EYATAECDNLVQLVTHDRAGEWVLEDLLVDAEQRLADEGIGGDIYLFKNN
**::.* : :: * *:*** *:* * :::*. . : *:***
Mflv_3075|M.gilvum_PYR-GCK VDGKGASYGTHENYLMSRQTPFSAVISGLTPFLVSRQVVTGSGRVGIGPS
Mvan_3451|M.vanbaalenii_PYR-1 VDGKGASYGSHENYLMSRQTPFSAIIAGLTPFLVSRQVVTGSGRVGIGPS
MSMEG_3897|M.smegmatis_MC2_155 VDGKGASYGSHENYLMSRQTPFSAVIAGLTPFMVSRQVVTGSGRVGIGPS
TH_0506|M.thermoresistible__bu VDGKGASYGSHENYLMSRQTPFSAVITGLIPFFVSRQVICGQGRVGIGPS
Mb2136c|M.bovis_AF2122/97 VDGKGASYGSHENYLMSRQTPFSAIITGLTPFLVSRQVVTGSGRVGIGPS
Rv2112c|M.tuberculosis_H37Rv VDGKGASYGSHENYLMSRQTPFSAIITGLTPFLVSRQVVTGSGRVGIGPS
MMAR_3087|M.marinum_M VDGKGASYGSHENYLMSRQTPFSAIIAGLTPFLVSRQVVTGSGRVGIGPS
MUL_2333|M.ulcerans_Agy99 VDGKGASYGSHENYLMSRQTPFSAIIAGLTPFLVSRQVVTGSGRVGIGPS
MLBr_01320|M.leprae_Br4923 VDGKGASYGAHENYLMSRQTPFSAIIAGLTPFLVSRQVVTGSGRVGIGPA
MAB_2170|M.abscessus_ATCC_1997 IDGKGASYGTHENYLMDRQTPFSAIIAGLTPFFVSRQVVTGSGRVGIGPS
MAV_2406|M.avium_104 TDSAGNSYGCHENYLIVRAGEFSRISDVLLPFLVTRQLICGAGKVLQTP-
*. * *** *****: * ** : * **:*:**:: * *:* *
Mflv_3075|M.gilvum_PYR-GCK GDEPGFQLSQRSDYIEVEVGLETTLKRGIINTRDEPHADADKYRRLHVII
Mvan_3451|M.vanbaalenii_PYR-1 GDEPGFQLSQRADYIEVEVGLETTLKRGIINTRDEPHADADKYRRLHVII
MSMEG_3897|M.smegmatis_MC2_155 GDEPGFQLSQRADYIEVEVGLETTLKRGIINTRDEPHADADKYRRLHVII
TH_0506|M.thermoresistible__bu GDEPGFQLAQRADYIEVEVGLETTLKRGIINTRDEPHADADKYRRLHVII
Mb2136c|M.bovis_AF2122/97 GDEPGFQLSQRSDYIEVEVGLETTLKRGIINTRDEPHADADRYRRLHVII
Rv2112c|M.tuberculosis_H37Rv GDEPGFQLSQRSDYIEVEVGLETTLKRGIINTRDEPHADADRYRRLHVII
MMAR_3087|M.marinum_M GDEPGFQLSQRSDYIEVEVGLETTLKRGIINTRDEPHADADRYRRLHVII
MUL_2333|M.ulcerans_Agy99 GDEPGFQLSQRSDYIEVEVGLETTLKRGIINTRDEPHADADRYRRLHVII
MLBr_01320|M.leprae_Br4923 GDEPGFQLSQRSDYIEVEVGLETTLKRGIINTRDEPHADADRYRRLHVIV
MAB_2170|M.abscessus_ATCC_1997 GDDAGFQLSQRADYIEVEVGLETTLKRGIINTRDEPHADADKYRRLHVII
MAV_2406|M.avium_104 -KAATFCLSQRAEHIWEGVSSATTRSRPIINTRDEPHADAEKYRRLHVIV
. . * *:**:::* *. ** .* ************::*******:
Mflv_3075|M.gilvum_PYR-GCK GDANLAETSTYLKVGATSLVLDLIEEGPQIGLELSDLTLARPVHAVHVVS
Mvan_3451|M.vanbaalenii_PYR-1 GDANLAETSTYLKLGTTSLVLDLIEEGPRIGADLSDLALARPVHAVHVLS
MSMEG_3897|M.smegmatis_MC2_155 GDANLAETSTYLKLGTTSLVLDLIEEG----VDLSDLALARPVHAVHVIS
TH_0506|M.thermoresistible__bu GDANLAETATYLKLGTTALVLDLIEEGHQIGIDLSDLTLARPVHAVHVIS
Mb2136c|M.bovis_AF2122/97 GDANLAETSTYLKLGTTALVLDLIEEGPAHAIDLTDLALARPVHAVHAIS
Rv2112c|M.tuberculosis_H37Rv GDANLAETSTYLKLGTTALVLDLIEEGPAHAIDLTDLALARPVHAVHAIS
MMAR_3087|M.marinum_M GDANLSETSTYLKLGTTALVLDLIEEGPAHGIDLTDLALARPVHAVHAIS
MUL_2333|M.ulcerans_Agy99 GDANLSETSTYLKLGTTALVLDLIEEGPAHGIDLTDLALARPVHAVHAIS
MLBr_01320|M.leprae_Br4923 GDANLAETSTYLKLGTTALVLDLIEEGPVHGIDLTDLTLARPVHAVHAIS
MAB_2170|M.abscessus_ATCC_1997 GDANLAETSTFLKVGTTSLVLDLIESG----VDLSDLALARPVHAVHVIS
MAV_2406|M.avium_104 GDSNMCETTTMLKVGTAALVLEMIEAG----VPFRDFSLDNPIRAIREVS
**:*:.**:* **:*:::***::** * : *::* .*::*:: :*
Mflv_3075|M.gilvum_PYR-GCK RDPLLRATVALADGREMTALALQRIYLDRVAKLVDSRDPDPRASHVLETW
Mvan_3451|M.vanbaalenii_PYR-1 RDPSLRATVALADGRELTGLALQRIYLDRVAKLVDSRDPDPRATHVLETW
MSMEG_3897|M.smegmatis_MC2_155 RDPSLRATVALADGRELTALALQRIYLDRVAKLVDSRDPDPRASHVIETW
TH_0506|M.thermoresistible__bu RDPTLRATVALADGRELTGLALQRIYLDRVAKLVDAKDPDPRASHILETW
Mb2136c|M.bovis_AF2122/97 RDPSLRATVALADGRELTGLALQRIYLDRVAKLVDSRDPDPRAADIVETW
Rv2112c|M.tuberculosis_H37Rv RDPSLRATVALADGRELTGLALQRIYLDRVAKLVDSRDPDPRAADIVETW
MMAR_3087|M.marinum_M RDPSLRTAVALLDGRELTGLALQRIYLERVAKLVDGRDPDPRASHVVETW
MUL_2333|M.ulcerans_Agy99 RDPSLRTAVALLDGRELTGFALQRIYLERVAKLVDGRDPDPRASHVVETW
MLBr_01320|M.leprae_Br4923 RDASLRATVTLVDGRELTGLALQRIYLDRVAKLVDSRDPDPRAADVVKTW
MAB_2170|M.abscessus_ATCC_1997 RDPSLRATVALADGRELTGLALQRIYLDRVAELVSSRDPDPQADQVLQVW
MAV_2406|M.avium_104 HDITGRRPVRLAGGRQASALDIQREYYSRAVEHLQTREPNAQIEQIVDLW
:* * .* * .**: :.: :** * .*..: :. ::*:.: .::. *
Mflv_3075|M.gilvum_PYR-GCK AHVLDLLE-RDPMECADLLDWPAKLRLLEGFRNRENLGWNAPRLHLVDLQ
Mvan_3451|M.vanbaalenii_PYR-1 AHILDLLE-RDPMECADLLDWPAKLRILEGFRHRENLGWNAPRLHLVDLQ
MSMEG_3897|M.smegmatis_MC2_155 ANVLDLLE-RDPMECAEILDWPAKLRLLEGFRQRENLTWQAPRLHLVDLQ
TH_0506|M.thermoresistible__bu AEVLDLLE-RDPMECAEILDWPAKLRLLEGFRQRENLSWNAPRLHLVDLQ
Mb2136c|M.bovis_AF2122/97 AHVLDQLE-RDPMDCAELLDWPAKLRLLDGFRQRENLSWSAPRLHLVDLQ
Rv2112c|M.tuberculosis_H37Rv AHVLDQLE-RDPMDCAELLDWPAKLRLLDGFRQRENLSWSAPRLHLVDLQ
MMAR_3087|M.marinum_M ADVLDRLE-RDPMDCAEVLDWPAKLRLLEGFRQRENLNWSAPRLHLVDLQ
MUL_2333|M.ulcerans_Agy99 ADVLDRLE-RDPMDCAEVLDWPAKLRLLEGFRQRENLNWSAPRLHLVDLQ
MLBr_01320|M.leprae_Br4923 VHVLDQLE-RDPMDCAELLDWPAKLRLLEGFRQRENLNWSAPRLHLVDLQ
MAB_2170|M.abscessus_ATCC_1997 AKVLDLLE-RDPMECADLLDWPAKLRLLEGFRQRENLSWSAPRLHLVDLQ
MAV_2406|M.avium_104 GRQLDAVESQDFAKVDTEIDWVIKRKLFQRYQDRYNMELSDPKIAQLDLA
** :* :* . :** * :::: ::.* *: . *:: :**
Mflv_3075|M.gilvum_PYR-GCK YSDVRLDKGLYNRLVARGSMKRLVTEQQVLDAVDNPPTDTRAYFRGECLR
Mvan_3451|M.vanbaalenii_PYR-1 YSDVRLDKGLYNRLVARGSMQRLVTEQQVLDAVENPPTDTRAYFRGECLR
MSMEG_3897|M.smegmatis_MC2_155 YSDVRLDKGLYNRLVARGSMKRLVTEQQVLDAVENPPTDTRAYFRGECLR
TH_0506|M.thermoresistible__bu YSDVRLDKGLYNRLVARGSMKRLVTEQQVLDAVDNPPPDTRAYFRGECLR
Mb2136c|M.bovis_AF2122/97 YSDVRLDKGLYNRLVARGSMKRLVTEHQVLSAVENPPTDTRAYFRGECLR
Rv2112c|M.tuberculosis_H37Rv YSDVRLDKGLYNRLVARGSMKRLVTEHQVLSAVENPPTDTRAYFRGECLR
MMAR_3087|M.marinum_M YSDVRLDKGLYNRLVARGSMKRLVSEHQVLEAVENPPTDTRAYFRGECLR
MUL_2333|M.ulcerans_Agy99 YSDVRLDKGLYNRLVARGSMKRLVSEHQVLDAVENPPTDTRAYFRGECLR
MLBr_01320|M.leprae_Br4923 YSDVRLDKGLYNRLVARGSMKRLVNEHQVLRAVNNPPTDTRAYFRGECLR
MAB_2170|M.abscessus_ATCC_1997 YSDVRLDKGLYNRLVARGSMQRMVTEQQVLDAVTRPPSDTRAYFRGECLR
MAV_2406|M.avium_104 YHDIKRGRGVFDLLQRKGLAARVTTDEDIAEAVDTPPQTTRARLRGEFIS
* *:: .:*::: * :* *:..:.:: ** ** *** :*** :
Mflv_3075|M.gilvum_PYR-GCK RFGADIAAASWDSVIFDLGGDSLVRIPTLEPLRGSKAHVGALLDSVDSAV
Mvan_3451|M.vanbaalenii_PYR-1 RFGADIAAASWDSVIFDLGGDSLVRIPTLEPLRGSKAHVGALLDSVDSAV
MSMEG_3897|M.smegmatis_MC2_155 RFGADIAAASWDSVIFDLGGDSLVRIPTLEPLRGSKAHVGALLDSVDSAV
TH_0506|M.thermoresistible__bu RFGSDIAAASWDSVIFDLGGDSLVRIPTLEPLRGSKAHVGELLDSVDSAV
Mb2136c|M.bovis_AF2122/97 RFGADIAAASWDSVIFDLGGDSLVRIPTLEPLRGSKAHVGALLDSVDSAV
Rv2112c|M.tuberculosis_H37Rv RFGADIAAASWDSVIFDLGGDSLVRIPTLEPLRGSKAHVGALLDSVDSAV
MMAR_3087|M.marinum_M RFGADIAAASWDSVIFDLGGDSLVRIPTLEPLRGSKAHVGALLDSVDSAV
MUL_2333|M.ulcerans_Agy99 RFGADIAAASWDSVIFDLGGDSLVRIPTLEPLRGSKAHVGALLDSVNSAV
MLBr_01320|M.leprae_Br4923 RFSADIAAASWDSVIFDLGGDSLVRIPTLEPLRGSKAHVGALLDSVDSAA
MAB_2170|M.abscessus_ATCC_1997 KFGADIAAASWDSVIFDLGGDSLVRIPTLEPLRGSKSHVGALLDSVDSAA
MAV_2406|M.avium_104 AAQAAGRDFTVDWVHLKLNDQAQRTVLCKDPFRAVDERVKRLIASM----
: : * * :.*..:: : :*:*. . :* *: *:
Mflv_3075|M.gilvum_PYR-GCK ELVEQLTN------------
Mvan_3451|M.vanbaalenii_PYR-1 ELVEQLTT------------
MSMEG_3897|M.smegmatis_MC2_155 ELVEQLTN------------
TH_0506|M.thermoresistible__bu ELVEQLTT------------
Mb2136c|M.bovis_AF2122/97 ELVEQLTAEPR---------
Rv2112c|M.tuberculosis_H37Rv ELVEQLTAEPR---------
MMAR_3087|M.marinum_M ELVEQLTS------------
MUL_2333|M.ulcerans_Agy99 ELVEQLTS------------
MLBr_01320|M.leprae_Br4923 ELVEQLTTKPVDPGKVGLDR
MAB_2170|M.abscessus_ATCC_1997 ELVDQLTT------------
MAV_2406|M.avium_104 --------------------