For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MTEQVARHRTAMTRAALSRPVALAISDGVLNSSLSVFDYGCGRGDDLRNLTALGFQTDGWDPSHRPEAAL RPADIVNIGYVVNVIDDRAERRETLQRAWNLAKQVLVVSARLVWEARDLEGRPYADGLVTRTGTFQKFYE QAELAAWIEESLGVKPVAAAPGIFYVFRDAARAHEFLATRAYTYRPRMRADPHAVYETHKQTLAPLLDFL SMHARSPRSGELEESAEVQIRDQFSSIASATNLIRQVTDDDYWDDVRLQRRQEMLVYIAMSRFGRRPRFS ELAKTLAADIKTHLGKYSDACLQADRLLLATGDPAIVLVSARSSSVGKQTPSALYVHRSALGHLPPVLRV YEACGRVLAGTVEHANMVKLSVKEPQVSYLTYPDFDRDPHPTLRSAVTVNLRRLSVDWRDYSRSENPPLL HRKEEFVGPDHPKRSLYERLTRAEVNAGLYAHPERIGTLIGWQSTMHAAGVSVRGHRLVLRQALSGTDAK
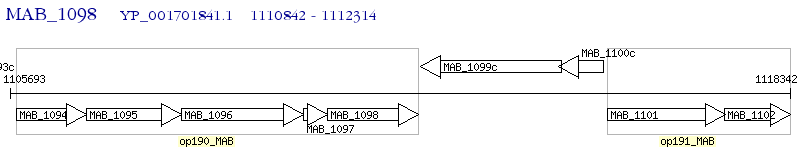
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. abscessus ATCC 19977 | MAB_1098 | - | - | 100% (490) | hypothetical protein MAB_1098 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | - | - | - | - | - |
| M. gilvum PYR-GCK | - | - | - | - | - |
| M. tuberculosis H37Rv | - | - | - | - | - |
| M. leprae Br4923 | - | - | - | - | - |
| M. marinum M | - | - | - | - | - |
| M. avium 104 | - | - | - | - | - |
| M. smegmatis MC2 155 | MSMEG_1246 | - | 3e-94 | 40.88% (477) | hypothetical protein MSMEG_1246 |
| M. thermoresistible (build 8) | - | - | - | - | - |
| M. ulcerans Agy99 | - | - | - | - | - |
| M. vanbaalenii PYR-1 | - | - | - | - | - |
CLUSTAL 2.0.9 multiple sequence alignment
MAB_1098|M.abscessus_ATCC_1997 MTE---QVARHRTAMTRAALSRPVALAISDGVLNSSLSVFDYGCGRGDDL
MSMEG_1246|M.smegmatis_MC2_155 MTVDEWRSARERTAIGRGDLSMPVRQCLRDEIISTKLSLLDYGCGRGQDV
** : **.***: *. ** ** .: * ::.:.**::*******:*:
MAB_1098|M.abscessus_ATCC_1997 RNLTALGFQTDGWDPSHRPEAALRPADIVNIGYVVNVIDDRAERRETLQR
MSMEG_1246|M.smegmatis_MC2_155 ARLSKMGVKAHGWDPHFAPDAPLVERDVVLMTYVLNVIENAQERRETLRK
.*: :*.::.**** . *:*.* *:* : **:***:: ******::
MAB_1098|M.abscessus_ATCC_1997 AWNLAKQVLVVSARLVWEARDLEGRPYADGLVTRTGTFQKFYEQAELAAW
MSMEG_1246|M.smegmatis_MC2_155 AWNLASKVLVVSCRLTWEQSSVRGAELGDGIVTSRNTFQHFFKPSEMREL
*****.:*****.**.** .:.* .**:** .***:*:: :*:
MAB_1098|M.abscessus_ATCC_1997 IEESLGVKPVAAAPGIFYVFRDAARAHEFLATRAYTYRPRMRADPHAVYE
MSMEG_1246|M.smegmatis_MC2_155 VEESTGRRCVSPAPGVVYAFRHDEDRFAYLARDAISDFEWTAS------E
:*** * : *:.***:.*.**. . :** * : : *
MAB_1098|M.abscessus_ATCC_1997 THKQTLAPLLDFLSMHARSPRSGELEESAEVQIRDQFSSIASATNLIRQV
MSMEG_1246|M.smegmatis_MC2_155 DYASAVAELVAFAEKRGRPPLFEEIPEQLLPQLG-KFS-RRSLLEVIQKG
: .::* *: * . :.*.* *: *. *: :** * ::*::
MAB_1098|M.abscessus_ATCC_1997 TDDDYWDDVRLQRRQEMLVYIAMSRFGRRPRFSELAKTLAADIKTHLGKY
MSMEG_1246|M.smegmatis_MC2_155 ASPDRVEEGRRRSTLDTLLYLGTSIFNGRASFSDLPPGIQADIKHCFKSY
:. * :: * : : *:*:. * *. *. **:*. : **** : .*
MAB_1098|M.abscessus_ATCC_1997 SDACLQADRLLLATGDPAIVLVSARSSSVGKQTPSALYVHRSALGHLPPV
MSMEG_1246|M.smegmatis_MC2_155 REACARADRLLAKIRDDTYIRG-AMQNSPGKLTATALYVHRRAVSKMPVV
:** :***** * : : * ..* ** *.:****** *:.::* *
MAB_1098|M.abscessus_ATCC_1997 LRVYEACGRVLAGTVEHANMVKLSVKEPQVSYLTYPDFDRDPHPTLRSAV
MSMEG_1246|M.smegmatis_MC2_155 LRLYEHCGFVAAGRPNDWNILKLDHRGRRVSWSSYPDFGTSPHPTLDWTY
**:** ** * ** :. *::**. : :**: :****. .***** :
MAB_1098|M.abscessus_ATCC_1997 TVNLRRLSVDWRDYSRSENPPLLHRKEEFVGPDHPKRSLYERLTRAEVNA
MSMEG_1246|M.smegmatis_MC2_155 GVDMLTLESAFQSFGSRENRPLLHRKEEFLDPDDPDVPKYRRLTTAEVRA
*:: *. ::.:. ** *********:.**.*. . *.*** ***.*
MAB_1098|M.abscessus_ATCC_1997 GLYAHPERIGTLIGWQSTMHAAGVSVRGHRLVLRQALSGTDAK
MSMEG_1246|M.smegmatis_MC2_155 GLYQNPTLIGLEKGWATELERCGVELRGHRLVKRKPS------
*** :* ** ** : :. .**.:****** *:.